Genomics
Accelerated Genomics
Speedup your genomics application from your browser using the power of accelerators
Speed up your applications
from your browser
InAccel’s Accelerated Genomics suite allows easy acceleration of genomics applications through an easy-to-use graphical interface.
Using a web-based portal researchers can utilize the power of the FPGAs to speedup their genomics applications like sequence alignment (i.e. Smith-Waterman).
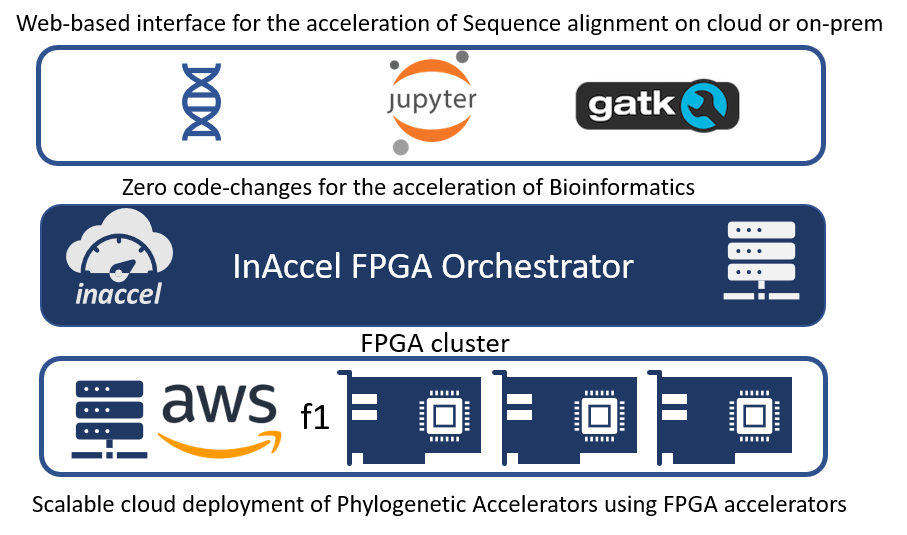
Accelerated genomics
InAccel’s Accelerated genomics studio allows easy to use framework for the utilization of FPGAs to speedup genomics applications like Smith Waterman and BWA MEM.
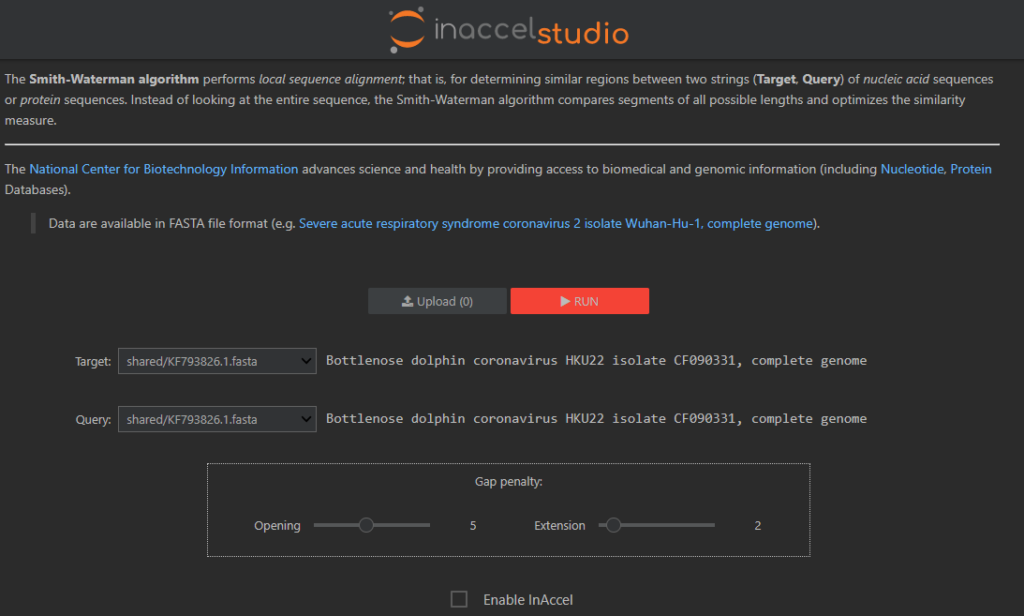
The web interface allows to upload your own data and to configure the required parameters.
It also allow to compare with CPU only execution and see instantly the power of FPGA accelerators.
Save cost from faster execution
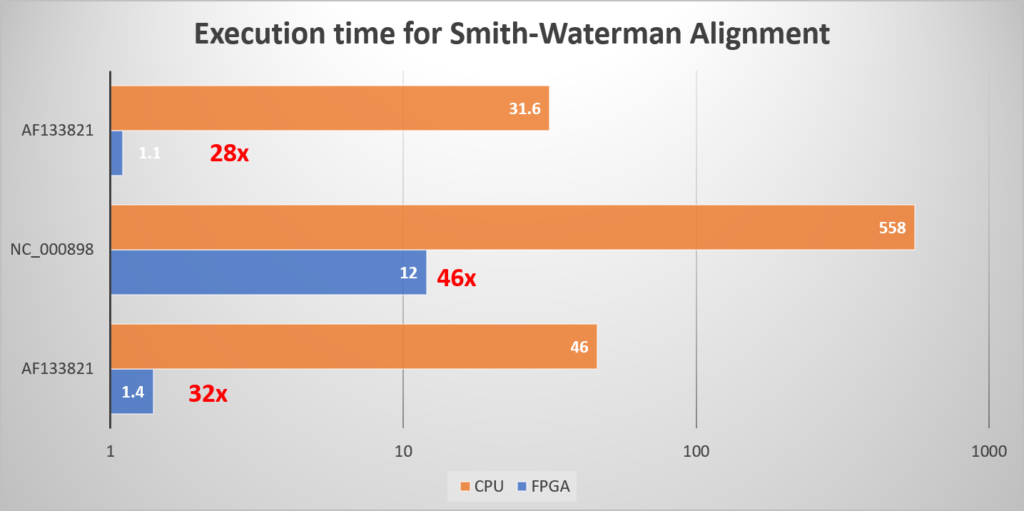
Depending on the dataset and the algorithm, the accelerated genomics platform can achieve up to 46 speedup compared to CPU-based execution.
At the same time, InAccel genomics solution can help reduce significantly the total cost of ownership as FPGA provide a more cost efficient solution based on CPUs and GPUs.
(The figure shows 8 applications in parallel deployed in 2 Intel PACs Arria 10 FPGAs vs 8 core CPU Xeon processor)
Note: The online web platform is available for demonstration purposes to show the easy of deployment using FPGA-based accelerators. Multiple users may share the available resources which may affect the performance of the applications. If you want to have exclusive access to speedup your applications contact us at info@inaccel.com.
